Research Profile
Over the past 20 years, research has shown that the intestinal microbiome, the community of microorganisms living in our gut, plays a crucial role in human health. Early studies in the mid‑2000s, enabled by new sequencing technologies, revealed that healthy people differ widely in their gut microbial composition and that certain microbial patterns (microbiome signatures) are linked to diseases. We understand now that the interplay between gut microbes and the body is complex and bidirectional, modulated by factors such as diet, metabolites, and antigens. These modulators and messengers can support health by regulating the immune system, intestinal barrier, and metabolic functions. However, when the functional balance between the microbiome and its host is disturbed, chronic inflammatory and neoplastic diseases can develop. CRC 1371 was launched in 2019 to move microbiome research beyond cataloging microbes and toward uncovering how they functionally contribute to inflammation and cancer. Over its planned 12-year duration, the consortium aims to identify disease-related microbial signatures, understand how they cause and contribute to disease, and translate this knowledge into targeted therapeutic strategies. Its interdisciplinary teams combine expertise in microbiology, immunology, gastroenterology, nutrition, and computational biology.
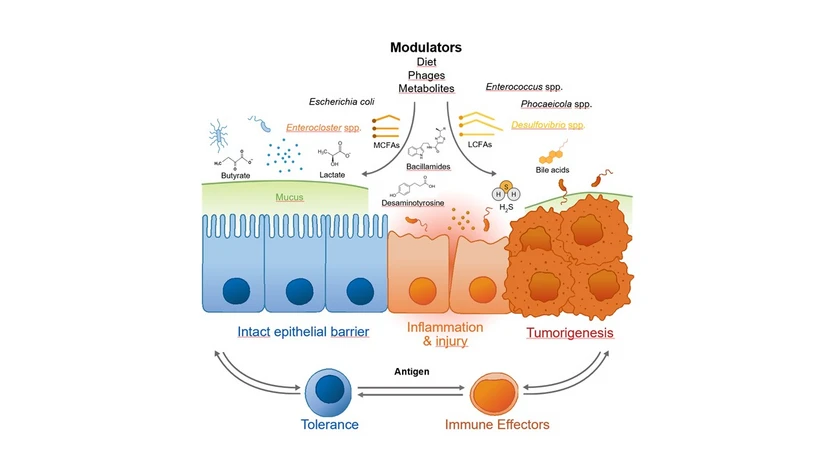
During the 1st funding phase, CRC 1371 built the infrastructure needed to study the microbiome at scale. It established standardized sampling protocols, unified computational pipelines, advanced experimental platforms, including gnotobiotic animals, and expanded patient access. These tools enabled the first proof-of-concept studies linking specific microbes and metabolites to inflammation and cancer development. The 2nd funding phase focused on refining the mechanisms and clinical relevance of microbiome signatures in defined disease endpoints, including infectious and immune-mediated inflammatory pathologies and cancer. Patients and models were implemented to test microbiome-based intervention strategies using diet, bacteriophages, and fecal microbiota transplantation. Cross-model and cross-cohort comparisons helped define the functional relevance and specificity of microbiome-based targets in disease. New technologies identified specific bacteria and metabolites that influence inflammation and therapeutic outcomes. Integrated analyses combining metabolomics, metagenomics, and clinical data supported biomarker discovery and translational progress, including early-stage therapeutic development.
CRC 1371 focusses on the analytical characterization (Pillar 1) oft he human microbiome and the successive functional evaluation (Pillar 2) of therapeutic interventions, which are based on the previous findeings. In particular, we apply three comprehensive strategies to characterize the functional and clinical relevance of the intestinal microbiome in disease models and patient cohorts:
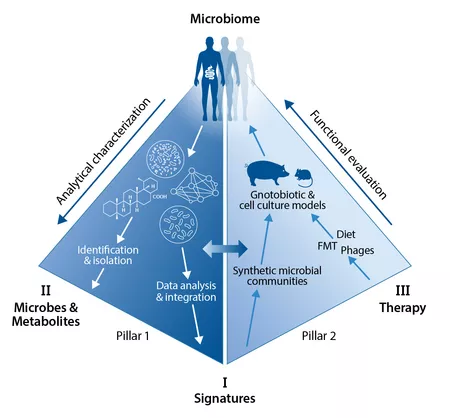
Strategy I - Characterization and functional evaluation of microbiome signatures in inflammation and cancer by applying genetically engineered and gnotobiotic animal models.
Strategy II - Identification and functional characterization of bioactive metabolites from the intestinal milieu using a comprehensive repertoire of analytical and computational methodologies.
Strategy III - Development and evaluation of microbiome-related therapeutic applications using fecal microbiota transplantation, bacteriophages and dietary interventions.